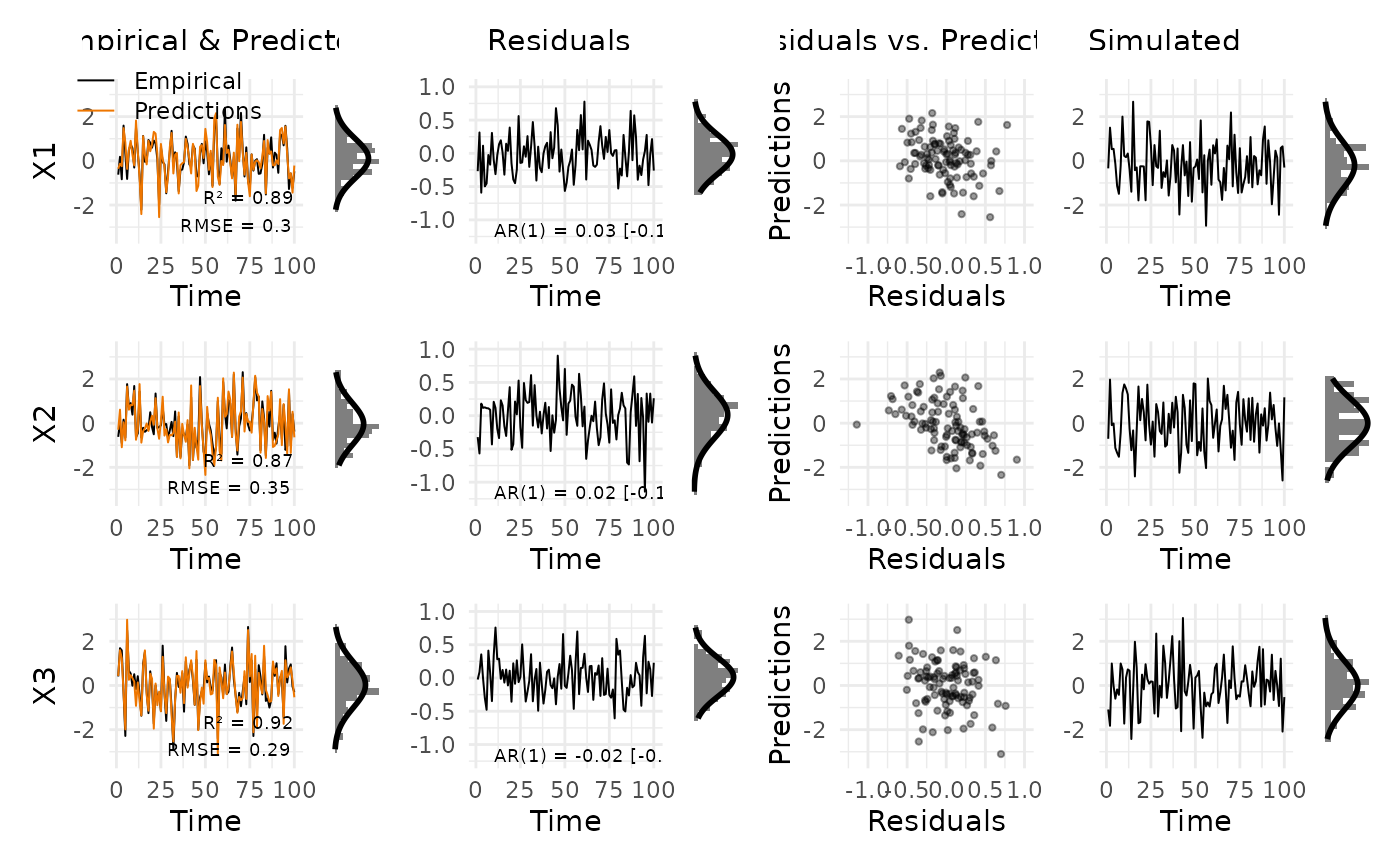
Creates a multi-panel diagnostic grid for a fitted VAR model. Each row corresponds to one variable; columns show (a) empirical data vs. predictions, (b) residuals over time, (c) residuals vs. predictions scatter, and (d) data simulated from the estimated model. Each time-series panel is accompanied by a marginal histogram with a Gaussian overlay.
Arguments
- data
A `var_data` object created with [new_var_data()].
- subject
Integer. Index of the subject to plot. Defaults to `1`.
- vars
Character or integer vector selecting variables to include. Defaults to all variables.
- panels
Character vector controlling which columns are shown. Any subset of `c("data", "residuals", "scatter", "simulated")`, in that order. Defaults to all four.
- colors
Named list controlling line colours. Recognised elements: `empirical` (default `"black"`) and `predicted` (default `"darkorange2"`). Partial lists are merged with the defaults.
- theme
A `ggplot2::theme()` object added on top of [theme_varcheck()]. Use this to override individual theme elements.
- ylim_data
Numeric vector of length 2. Shared y-limits for the data-scale panels (empirical/predicted/simulated). Auto-computed from the data if `NULL`.
- ylim_res
Numeric vector of length 2. Shared y-limits for the residual panels. Auto-computed from the residuals if `NULL`.